It is key to the process of protein synthesis taking place at ribosomes. What is this organelle?
How does the jail cell convert Deoxyribonucleic acid into working proteins? The process of translation tin can be seen every bit the decoding of instructions for making proteins, involving mRNA in transcription as well as tRNA.
The genes in DNA encode poly peptide molecules, which are the "workhorses" of the prison cell, conveying out all the functions necessary for life. For example, enzymes, including those that metabolize nutrients and synthesize new cellular constituents, as well equally Deoxyribonucleic acid polymerases and other enzymes that brand copies of Dna during cell partition, are all proteins.
In the simplest sense, expressing a factor means manufacturing its corresponding protein, and this multilayered process has ii major steps. In the first pace, the information in Dna is transferred to a messenger RNA (mRNA) molecule by way of a process called transcription. During transcription, the Dna of a cistron serves as a template for complementary base-pairing, and an enzyme chosen RNA polymerase Ii catalyzes the formation of a pre-mRNA molecule, which is and then processed to form mature mRNA (Figure 1). The resulting mRNA is a single-stranded copy of the cistron, which adjacent must be translated into a protein molecule.
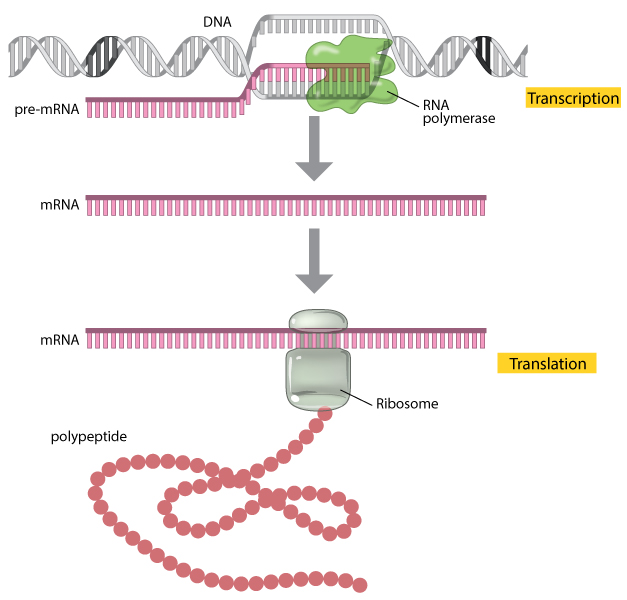
Figure 1: A gene is expressed through the processes of transcription and translation.
During transcription, the enzyme RNA polymerase (green) uses Dna equally a template to produce a pre-mRNA transcript (pinkish). The pre-mRNA is processed to class a mature mRNA molecule that can be translated to build the poly peptide molecule (polypeptide) encoded by the original gene.
During translation, which is the second major stride in gene expression, the mRNA is "read" according to the genetic code, which relates the DNA sequence to the amino acid sequence in proteins (Figure two). Each grouping of three bases in mRNA constitutes a codon, and each codon specifies a item amino acid (hence, it is a triplet lawmaking). The mRNA sequence is thus used equally a template to assemble—in order—the concatenation of amino acids that form a poly peptide.

Figure 2: The amino acids specified past each mRNA codon. Multiple codons can lawmaking for the same amino acid.
The codons are written v' to 3', equally they announced in the mRNA. AUG is an initiation codon; UAA, UAG, and UGA are termination (end) codons.
But where does translation have identify within a cell? What individual substeps are a part of this process? And does translation differ between prokaryotes and eukaryotes? The answers to questions such as these reveal a great deal almost the essential similarities between all species.
Where Translation Occurs
Inside all cells, the translation machinery resides inside a specialized organelle chosen the ribosome. In eukaryotes, mature mRNA molecules must go out the nucleus and travel to the cytoplasm, where the ribosomes are located. On the other hand, in prokaryotic organisms, ribosomes can attach to mRNA while it is still beingness transcribed. In this situation, translation begins at the 5' end of the mRNA while the 3' terminate is still attached to DNA.
In all types of cells, the ribosome is composed of ii subunits: the large (50S) subunit and the small-scale (30S) subunit (S, for svedberg unit, is a measure of sedimentation velocity and, therefore, mass). Each subunit exists separately in the cytoplasm, but the 2 bring together together on the mRNA molecule. The ribosomal subunits contain proteins and specialized RNA molecules—specifically, ribosomal RNA (rRNA) and transfer RNA (tRNA). The tRNA molecules are adaptor molecules—they have one end that can read the triplet code in the mRNA through complementary base-pairing, and another terminate that attaches to a specific amino acid (Chapeville et al., 1962; Grunberger et al., 1969). The idea that tRNA was an adaptor molecule was offset proposed by Francis Crick, co-discoverer of Dna construction, who did much of the central work in deciphering the genetic lawmaking (Crick, 1958).
Within the ribosome, the mRNA and aminoacyl-tRNA complexes are held together closely, which facilitates base of operations-pairing. The rRNA catalyzes the zipper of each new amino acrid to the growing chain.
The Offset of mRNA Is Not Translated
Interestingly, not all regions of an mRNA molecule stand for to detail amino acids. In particular, there is an area near the v' end of the molecule that is known every bit the untranslated region (UTR) or leader sequence. This portion of mRNA is located betwixt the kickoff nucleotide that is transcribed and the start codon (AUG) of the coding region, and it does not affect the sequence of amino acids in a poly peptide (Figure 3).
So, what is the purpose of the UTR? It turns out that the leader sequence is important because it contains a ribosome-binding site. In leaner, this site is known as the Smooth-Dalgarno box (AGGAGG), after scientists John Smooth and Lynn Dalgarno, who kickoff characterized information technology. A similar site in vertebrates was characterized by Marilyn Kozak and is thus known as the Kozak box. In bacterial mRNA, the v' UTR is normally brusk; in human mRNA, the median length of the 5' UTR is about 170 nucleotides. If the leader is long, it may contain regulatory sequences, including binding sites for proteins, that can affect the stability of the mRNA or the efficiency of its translation.
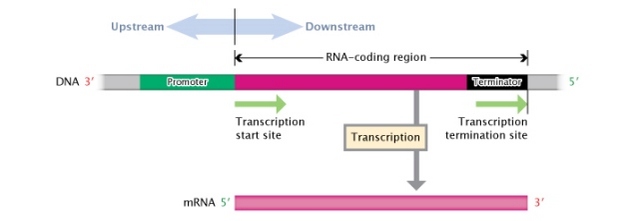
Figure 3: A Dna transcription unit.
A Deoxyribonucleic acid transcription unit is composed, from its 3' to 5' end, of an RNA-coding region (pink rectangle) flanked by a promoter region (green rectangle) and a terminator region (black rectangle). Regions to the left, or moving towards the 3' end, of the transcription start site are considered \"upstream;\" regions to the right, or moving towards the 5' end, of the transcription start site are considered \"downstream.\"
© 2014 Nature Education Adapted from Pierce, Benjamin. Genetics: A Conceptual Arroyo, second ed. All rights reserved. ![]()
Translation Begins Afterward the Assembly of a Circuitous Structure
The translation of mRNA begins with the formation of a circuitous on the mRNA (Effigy 4). First, 3 initiation cistron proteins (known equally IF1, IF2, and IF3) bind to the small subunit of the ribosome. This preinitiation complex and a methionine-carrying tRNA then bind to the mRNA, near the AUG start codon, forming the initiation circuitous.
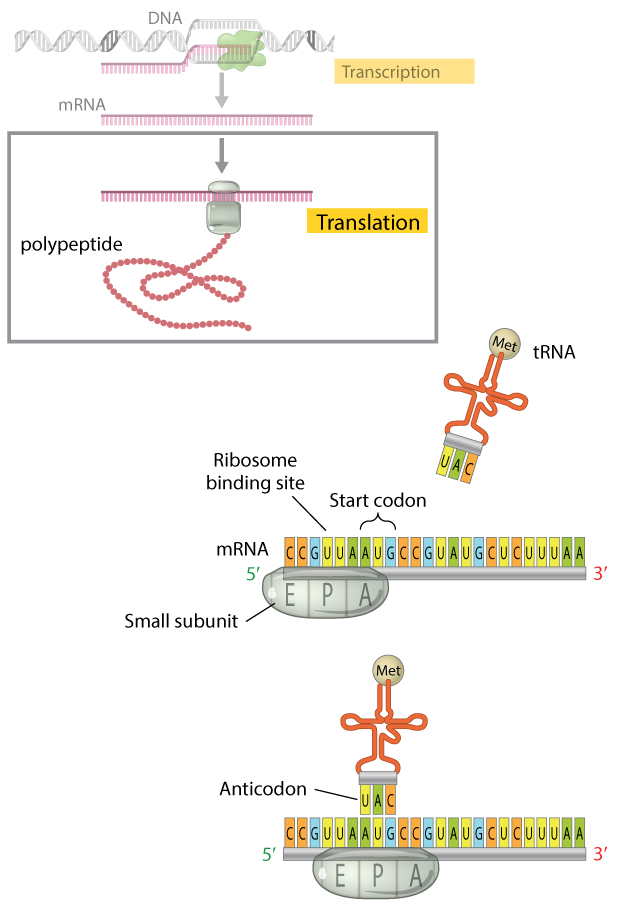
Figure 4: The translation initiation circuitous.
When translation begins, the small subunit of the ribosome and an initiator tRNA molecule gather on the mRNA transcript. The small subunit of the ribosome has three bounden sites: an amino acrid site (A), a polypeptide site (P), and an exit site (E). The initiator tRNA molecule carrying the amino acrid methionine binds to the AUG start codon of the mRNA transcript at the ribosome's P site where information technology will become the first amino acid incorporated into the growing polypeptide concatenation. Here, the initiator tRNA molecule is shown binding after the small ribosomal subunit has assembled on the mRNA; the lodge in which this occurs is unique to prokaryotic cells. In eukaryotes, the costless initiator tRNA offset binds the small-scale ribosomal subunit to course a complex. The circuitous then binds the mRNA transcript, so that the tRNA and the pocket-sized ribosomal subunit bind the mRNA simultaneously.
Although methionine (Met) is the first amino acid incorporated into any new poly peptide, information technology is not always the commencement amino acrid in mature proteins—in many proteins, methionine is removed after translation. In fact, if a large number of proteins are sequenced and compared with their known factor sequences, methionine (or formylmethionine) occurs at the Due north-terminus of all of them. However, not all amino acids are as likely to occur 2d in the chain, and the second amino acrid influences whether the initial methionine is enzymatically removed. For example, many proteins begin with methionine followed by alanine. In both prokaryotes and eukaryotes, these proteins have the methionine removed, so that alanine becomes the N-terminal amino acid (Table one). Withal, if the second amino acid is lysine, which is also oft the case, methionine is not removed (at least in the sample proteins that have been studied thus far). These proteins therefore begin with methionine followed by lysine (Flinta et al., 1986).
Table ane shows the N-terminal sequences of proteins in prokaryotes and eukaryotes, based on a sample of 170 prokaryotic and 120 eukaryotic proteins (Flinta et al., 1986). In the table, M represents methionine, A represents alanine, K represents lysine, South represents serine, and T represents threonine.
Table one: Northward-Terminal Sequences of Proteins
| Due north-Terminal Sequence | Per centum of Prokaryotic Proteins with This Sequence | Percent of Eukaryotic Proteins with This Sequence |
| MA* | 28.24% | 19.17% |
| MK** | ten.59% | 2.50% |
| MS* | 9.41% | xi.67% |
| MT* | vii.65% | 6.67% |
* Methionine was removed in all of these proteins
** Methionine was not removed from whatsoever of these proteins
Once the initiation complex is formed on the mRNA, the big ribosomal subunit binds to this complex, which causes the release of IFs (initiation factors). The big subunit of the ribosome has iii sites at which tRNA molecules can demark. The A (amino acid) site is the location at which the aminoacyl-tRNA anticodon base of operations pairs upward with the mRNA codon, ensuring that right amino acrid is added to the growing polypeptide chain. The P (polypeptide) site is the location at which the amino acid is transferred from its tRNA to the growing polypeptide chain. Finally, the E (exit) site is the location at which the "empty" tRNA sits earlier existence released dorsum into the cytoplasm to bind another amino acid and repeat the process. The initiator methionine tRNA is the only aminoacyl-tRNA that can bind in the P site of the ribosome, and the A site is aligned with the 2nd mRNA codon. The ribosome is thus ready to demark the second aminoacyl-tRNA at the A site, which volition be joined to the initiator methionine by the outset peptide bond (Figure 5).

Figure 5: The large ribosomal subunit binds to the minor ribosomal subunit to complete the initiation complex.
The initiator tRNA molecule, carrying the methionine amino acid that will serve as the kickoff amino acid of the polypeptide concatenation, is bound to the P site on the ribosome. The A site is aligned with the next codon, which volition be leap past the anticodon of the side by side incoming tRNA.
The Elongation Phase
The next phase in translation is known every bit the elongation stage (Figure 6). Commencement, the ribosome moves along the mRNA in the 5'-to-3'management, which requires the elongation gene Thousand, in a process called translocation. The tRNA that corresponds to the second codon can so demark to the A site, a step that requires elongation factors (in E. coli, these are called EF-Tu and EF-Ts), too as guanosine triphosphate (GTP) as an energy source for the process. Upon bounden of the tRNA-amino acid complex in the A site, GTP is broken to grade guanosine diphosphate (Gdp), then released along with EF-Tu to exist recycled by EF-Ts for the adjacent round.
Next, peptide bonds between the now-adjacent first and second amino acids are formed through a peptidyl transferase activity. For many years, it was thought that an enzyme catalyzed this step, simply recent show indicates that the transferase activity is a catalytic function of rRNA (Pierce, 2000). Later the peptide bond is formed, the ribosome shifts, or translocates, once again, thus causing the tRNA to occupy the Due east site. The tRNA is then released to the cytoplasm to pick up some other amino acid. In improver, the A site is now empty and set up to receive the tRNA for the next codon.
This procedure is repeated until all the codons in the mRNA have been read by tRNA molecules, and the amino acids attached to the tRNAs have been linked together in the growing polypeptide chain in the advisable order. At this signal, translation must be terminated, and the nascent protein must be released from the mRNA and ribosome.
Termination of Translation
There are three termination codons that are employed at the end of a protein-coding sequence in mRNA: UAA, UAG, and UGA. No tRNAs recognize these codons. Thus, in the place of these tRNAs, i of several proteins, chosen release factors, binds and facilitates release of the mRNA from the ribosome and subsequent dissociation of the ribosome.
Comparing Eukaryotic and Prokaryotic Translation
The translation procedure is very similar in prokaryotes and eukaryotes. Although different elongation, initiation, and termination factors are used, the genetic code is more often than not identical. As previously noted, in bacteria, transcription and translation take place simultaneously, and mRNAs are relatively short-lived. In eukaryotes, however, mRNAs have highly variable half-lives, are subject to modifications, and must exit the nucleus to be translated; these multiple steps offer boosted opportunities to regulate levels of poly peptide product, and thereby fine-tune factor expression.
References and Recommended Reading
Chapeville, F., et al. On the part of soluble ribonucleic acid in coding for amino acids. Proceedings of the National University of Sciences 48, 1086–1092 (1962)
Crick, F. On poly peptide synthesis. Symposia of the Society for Experimental Biology 12, 138–163 (1958)
Flinta, C., et al. Sequence determinants of Northward-terminal protein processing. European Journal of Biochemistry 154, 193–196 (1986)
Grunberger, D., et al. Codon recognition past enzymatically mischarged valine transfer ribonucleic acid. Science 166, 1635–1637 (1969) doi:10.1126/science.166.3913.1635
Kozak, Thousand. Betoken mutations close to the AUG initiator codon bear on the efficiency of translation of rat preproinsulin in vivo. Nature 308, 241–246 (1984) doi:10.1038308241a0 (link to commodity)
---. Signal mutations define a sequence flanking the AUG initiator codon that modulates translation by eukaryotic ribosomes. Jail cell 44, 283–292 (1986)
---. An analysis of 5'-noncoding sequences from 699 vertebrate messenger RNAs. Nucleic Acids Research 15, 8125–8148 (1987)
Pierce, B. A. Genetics: A conceptual approach (New York, Freeman, 2000)
Shine, J., & Dalgarno, L. Determinant of gene specificity in bacterial ribosomes. Nature 254, 34–38 (1975) doi:10.1038/254034a0 (link to article)
szymanskifrooking.blogspot.com
Source: https://www.nature.com/scitable/topicpage/translation-dna-to-mrna-to-protein-393/
0 Response to "It is key to the process of protein synthesis taking place at ribosomes. What is this organelle?"
Post a Comment